Two books launched in June have systematized an alternative method for classifying organisms based on their evolutionary history and relationships to ancestors and descendants, regardless of their anatomical characteristics. It does not use the traditional hierarchical taxonomic categories, such as domains, kingdoms, phyla, classes, orders, families, and genders. These divisions derive from the ideas presented in the mid-eighteenth century in successive editions of the famous work Systema Naturae, by the Swedish naturalist Carl von Linnaeus (1707–1778). Defenders of PhyloCode, the formal name of the recently proposed initiative, consider the Linnaean standard categories—still one of the pillars of modern taxonomy—as abstractions detached from biological reality that no longer make sense in the face of advancements in phylogeny. They also criticize the difficulties of incorporating the discovery of new species, and revising their relatedness to other organisms, in a system based on impenetrable taxonomic divisions.
American biologist Philip Cantino of Ohio University is one of the creators of PhyloCode, with colleague Kevin de Queiroz at the National Museum of Natural History in Washington, D.C. Interviewed by email, Cantino told Pesquisa FAPESP, “Linnaeus’s contribution to biology was revolutionary and remarkably long-lasting, but it was made pre-evolution and predates the publication of Charles Darwin’s The Origin of Species by a century.” Their book, titled International Code of Phylogenetic Nomenclature (PhyloCode), contains the fundamental rules and guidelines of their proposal. “Despite the almost universal acceptance of the premise that classification should be based on phylogenetic relationships, biologists continue to name clades [groups of organisms descended from a common ancestor] using a pre-evolutionary system.” Darwin’s theory of evolution proposed the idea, now widely accepted, that contemporary species descend from past life forms.
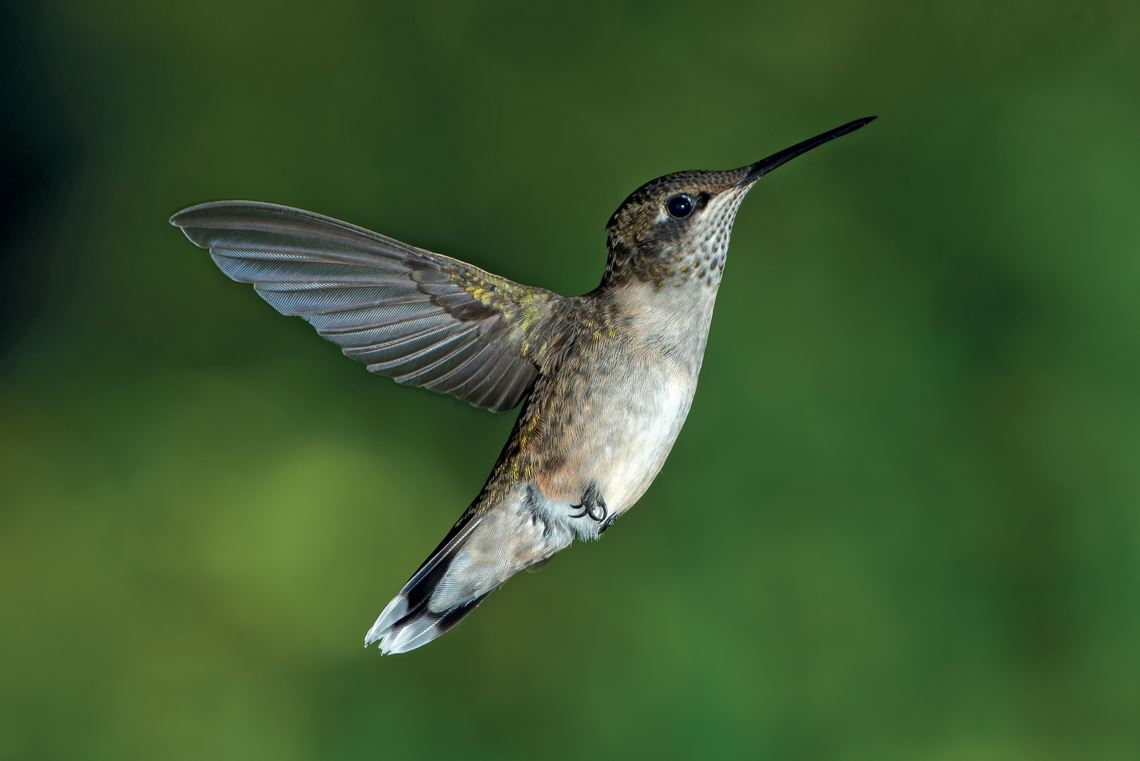

Pslawinski / Wikimedia Commons
Hummingbird: phylogenetic studies indicate that birds descended from dinosaursPslawinski / Wikimedia CommonsThe volume Phylonyms: A Companion to the PhyloCode, organized by de Queiroz, Cantino, and American paleontologist Jacques Gauthier, from Yale University, includes the definitions, based on the PhyloCode, of about 300 life-form lineages, known as clades, from microorganisms to plants and animals. Among the hundreds of collaborators on Phylonyms are three researchers from Brazilian institutions who were the main authors of the definitions for certain groups of organisms. Paleontologist Max Langer, from the University of São Paulo (USP), Ribeirão Preto campus, coordinated development of the text on dinosaurs and also wrote those on the Saurischia and Sauropodomorpha clades. In partnership with Richard Olmstead of the University of Washington, botanist Lúcia Lohmann, from USP produced the entry on Bignoniaceae, the group of plants that includes trees, shrubs, and lianas. Zoologist Martin Lindsey Christoffersen, from the Federal University of Paraíba (UFPB), wrote the definitions for two clades of decapod crustaceans, Polycarpidea and Prochelata.
Researchers in some areas of science, such as vertebrate paleontology and botany, have long adopted PhyloCode in their scientific works. But the proposal is not recognized by the international nomenclature codes and is still far from achieving consensus in academia. In this new classification system, there are only two categories, species and clades. PhyloCode’s originators thought about changing the method of naming species, but they gave up on the idea. “That would cause a lot of problems,” de Queiroz recognizes. Since the publication of Systema Naturae in 1758, species have been named according to the precepts of the binomial nomenclature system. Each unique form of life is designated by two words, usually of Latin (or Greek) origin, written in italics. The first, which is less specific, defines the genus, and its first letter is capitalized. The second, which may be derived from a specific characteristic of the organism in question, its geographical distribution, or simply a reference (as an homage) to a person, determines the species itself. This second word is written in all lowercase characters. Following this standard, Linnaeus named the species of modern man Homo sapiens. In Latin, homo means to be human and sapiens, wise or intelligent.
The concept of clade, a pillar of PhyloCode, is associated with the evolutionary idea that life forms descend from their predecessors, from which they inherit certain characteristics. The most commonly used metaphor is the tree of life. The base of its trunk represents the appearance of life on Earth approximately 4 billion years ago, beginning with a hypothetical population of microorganisms that ultimately all creatures—past and present—derive from. With the passage of time the tree of life generates branches that subdivide into other branches, and so on. At a certain point, some of these nodes develop in parallel, independently, and others remain connected. Still others are even cut off (when a species goes extinct). Stemming from Greek roots, the word “clade” means branch. In biology, in evolutionary terms, a clade (also called a monophyletic group) is made up of all the organisms that descend from a specific common ancestor.
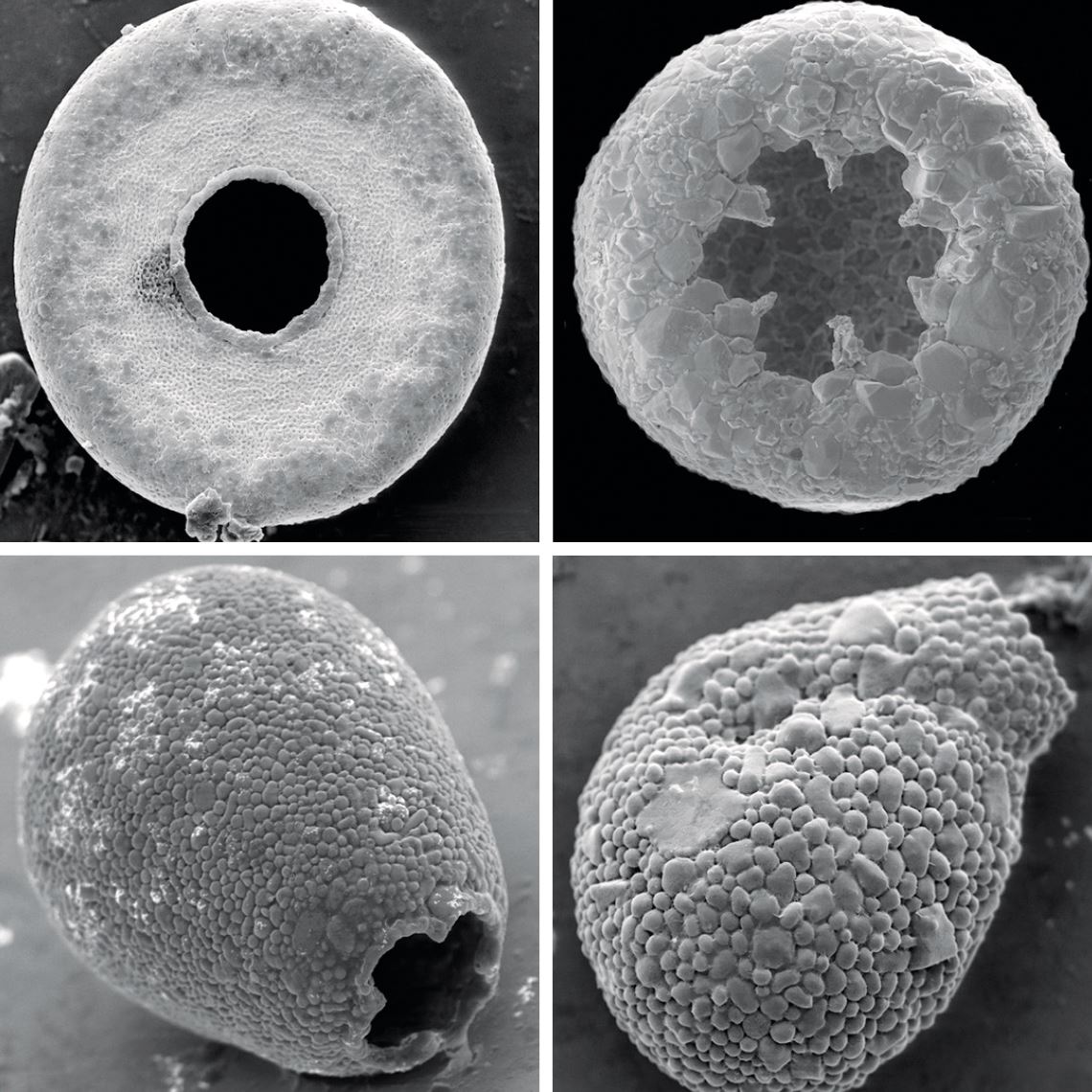
Daniel Lahr
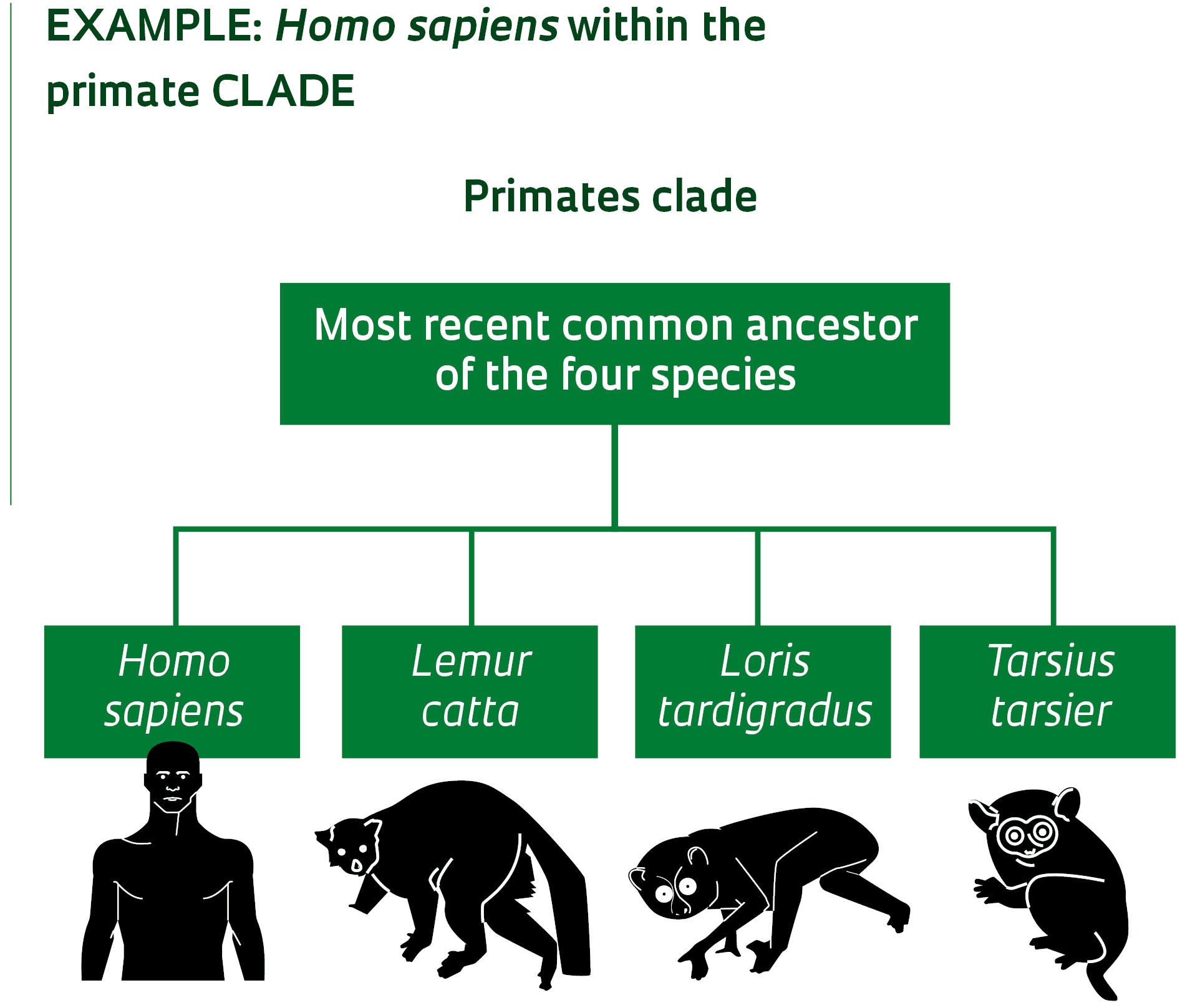
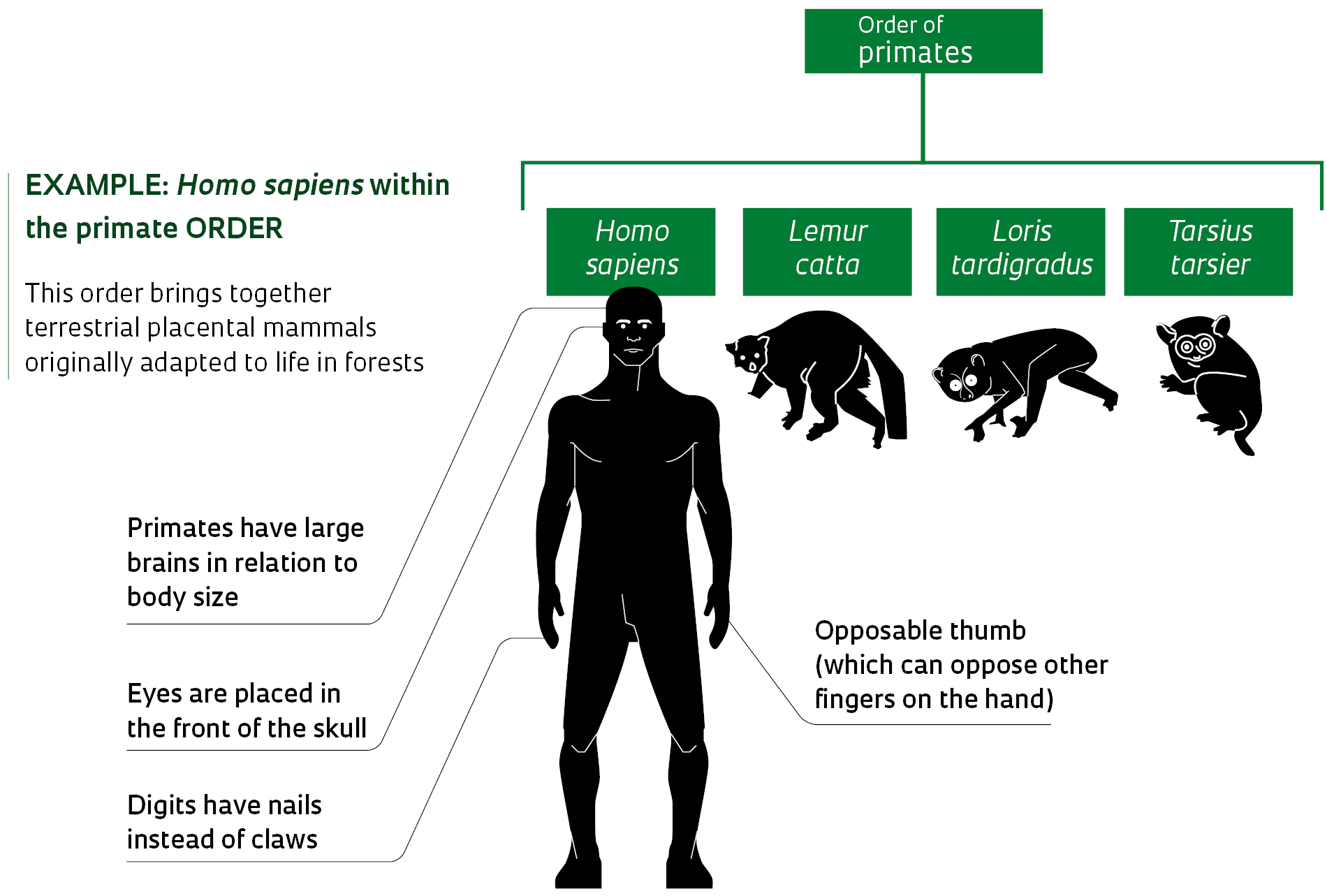
Microorganisms, such as these shelled amoebas from the group Arcellinida, can be difficult to classify without genetic dataDaniel LahrSound complicated? Examples help with understanding the concept. In traditional classification, primates form an order, a taxonomic level that can encompass numerous subdivisions, such as suborder, superfamily, family, subfamily, tribe, genus, and species. Roughly speaking, the order of primates is made up of lemurs, lorises, tarsiers, monkeys, and modern man. In PhyloCode, primates form one clade, defined as: the most recent common ancestor of the species Lemur catta (ring-tailed lemur), Loris tardigradus (red lory), Tarsius tarsier (tarsier) and Homo sapiens (modern man) and all its descendants. In practice, the life forms that belong to the order of primates, in the context of traditional taxonomy, and to the primate clade, from the perspective of PhyloCode, are essentially the same. This is because the primate clade was defined based on the ancestral relationships of representatives of the four main groups traditionally housed in the primate order (a lemur, a loris, a tarsier, and modern man).
But the reasoning that governs the constitution of clades in PhyloCode is different from that of the various taxonomic categories in the rank-based classifications. “The Linnaean system embodies a static and typological world view,” explains Christoffersen. “It is based on ‘essences’ [marked characteristics] to define taxa [groups of organisms or populations that form a unit] and type specimens to characterize these essences. With the theory of evolution, it is inevitable that this view will be replaced by one of the world in transformation.” (Type specimens are the individual specimen used as a reference to describe a species of organism.)
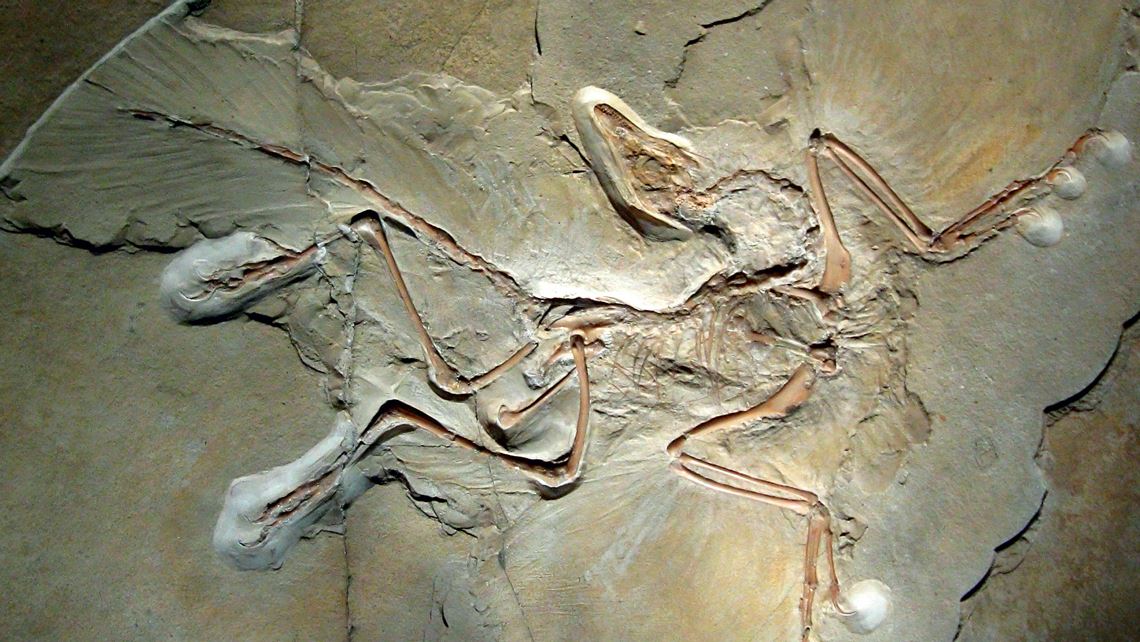
Phylogeny as a basis for classifying species can lead to conclusions that sound strange to laypeople, although they are now widely accepted. The case of birds is perhaps the most emblematic. There is virtually a consensus today among scientists that this group of animals is the only one that includes the living descendants of dinosaurs, which are considered to have gone extinct 65 million years ago. The discovery of half-meter-long dinosaur fossils with feathers and wings that lived at least 150 million years ago, such as the famous specimens from the Archeopteryx genus, support this conclusion. These modern bird ancestors are classified as members of the theropod group, usually composed of bipedal carnivores. Theropods form one of the three component lineages of dinosaurs, alongside sauropodomorphs (herbivores, usually quadrupeds, large and with elongated necks) and ornithischians (species with horns, armor, or beaks similar to those of ducks).
It’s no coincidence that the PhyloCode definition of the Dinosauria clade, the dinosaurs, is based on the relationship of known representatives of these three lineages: the ornithischian Iguanodon bernissartensis, the theropod Megalosaurus bucklandii and the sauropod Cetiosaurus oxoniensis. According to this new classification system, all descendants of the last common ancestor of these three species should be considered dinosaurs. “We’re not concerned with finding anatomical features that are exclusive or that define what a dinosaur was,” explains Langer, author of the Phylonyms entry on these long-extinct creatures. “What matters is whether the evolutionary history of a fossil puts it inside or outside the clade.” Currently, this task is done with the aid of computer programs that cross-reference mountains of data regarding anatomical characteristics (such as if they have feathers, are biped, or quadruped) and genetic material (if it exists) from the species under analysis. They then provide probable evolutionary trees in which a species or set of species may be included within (or excluded from) a clade.
Although he doesn’t consider PhyloCode the solution to all the problems in taxonomy, evolutionary microbiologist Daniel Lahr from USP’s Biosciences Institute (IB) believes that this type of classification can be useful for researchers focused on the study of certain groups of organisms. “For the moment it’s the best approach we have, specifically for those working with species that are defined based on a set of genetic and fossil data, and not as much by their morphology,” comments Lahr. “But certain well-established areas within zoology and botany should resist taking the cladistic approach.” In early 2019, the researcher published an article in the journal Current Biology in which he reconstructed the evolutionary history of a group of shelled amoebas, the Arcellinida, which emerged about 750 million years ago. Using mathematical algorithms and DNA analysis of amoebae from this group that exist in nature today, he created a phylogeny for this clade, using notions both from PhyloCode and more traditional taxonomy.
There is presently no unified standard that is used in all branches of biology for naming and establishing taxonomic categories. There is one nomenclature code for plants, algae, and fungi, another for animals, a third for bacteria and archaea, and a fourth for viruses. Each code determines how many taxonomic levels can be used to classify its groups of organisms. Botany, for example, provides 24 categories from kingdom to subform, while viral biology has 15. “We need to abandon this provincialism in each area, adopt one code, and move towards a single system,” the IB researcher proposes. It’s difficult to predict whether PhyloCode will spread throughout biology or remain an approach whose use is restricted to certain niches of researchers. Despite its limitations, Linnaean-based taxonomy is still useful and widely adopted.
- Groups organisms by their phylogeny. Their evolutionary history determines which clade, and which branch of the tree of life each species belongs to
- A clade is the set of organisms or species that descend from their most recent common ancestor
- There are no hierarchical categories, such as kingdom, order, family, and genus
- There are species and clades
- A phylogeny is composed of more inclusive clades that contain less comprehensive clades
The Phylonyms book defines this clade as the group of organisms that descend from the most recent common ancestor of four species: Homo sapiens (modern man), Lemur catta (ring-tailed lemur), Loris tardigradus (red slender loris) and Tarsius tarsier (tarsier). In this case, modern man not only belongs to the primate clade, but is also one element of the very definition of this group of organisms
- Organizes living things based on their natural characteristics, commonly their morphology (external appearance)
- Adopts a hierarchical system of taxonomic categories, such as dominion, kingdom, phylum, class, order, family, genus, species, and others
- The four major international nomenclature codes that classify different life forms—the plants, algae and fungi; animals; bacteria and archaea; and viruses—are based on hierarchical systems