
RCPedia
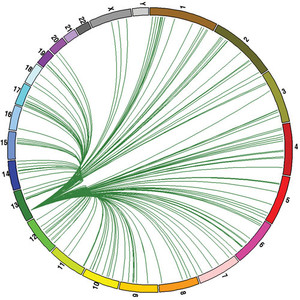
Genes in movement: The figure above relates the points of origin and destination of the 8,000 retrocopies in the 24 human chromosomes. Each colored line represents one gene, copied in its original chromosome of the same color. The copies – or retrocopies – scatter and establish themselves in other differently colored chromosomesRCPediaGenes known as retrocopies, which generate copies of themselves and insert themselves into other genes, likely contributed to the appearance of new species of primates from common ancestors, as well as to differentiation between individuals of each species. The results of a study by Brazilian and U.S. geneticists who compared the genome of six species of primates conclude that this type of gene plays an enhanced role in the adaptation and evolution of living organisms. More broadly speaking, retrocopies are no longer viewed as exotic or useless, as they were until just a few years ago. Instead, they are shown to be an important mechanism of renewal and regulation of the genome, the set of genes in an organism. The dozens or hundreds of copies established in various chromosomes, the structures that house the genes, are now being viewed as a possible mechanism for safeguarding essential information for survival.
“Some retrocopies are like gene fossils that can contain strands of DNA that were active at the time they emerged among the primates, nearly 60 million years ago, but are no longer so,” explains molecular scientist Pedro Galante, coordinator of the bioinformatics laboratory of the Teaching and Research Institute at the Syrian-Lebanese Hospital in São Paulo, where he works on formulating and verifying hypotheses about the possible origins and functions of retrocopies in the human genome.

L_DE_G/ FLICKERA marmoset, one of the species of primates whose genome was examinedL_DE_G/ FLICKER
The conclusions are now being discussed calmly, such as at an international seminar on genomics and cancer held in June 2016 in São Paulo, but that has not always been the case. In the 1940s, U.S. geneticist Barbara McClintock caused an uproar when she showed that mobile elements of the genome then called jumping genes, a concept inconceivable at the time, were responsible for the color variation among kernels of corn.
By comparing the genome of primates, Galante and Fábio Navarro, currently a postdoctoral researcher at Yale University in the United States, have determined that the retrocopies account for 45% of the genome of humans, chimpanzees and gorillas. They then concluded that nearly half of these mobile elements found in the six species examined must have originated during the early days of the evolution of primates, which began to form their own species nearly 90 million years ago.
Galante and Navarro identified 154 common and potentially functional retrocopies among the six species, which is not surprising since they are of the same evolutionary lineage. More interesting was seeing the unique retrocopies of each species, although their functions are still little known. Humans have 127 unique retrocopies; gorillas have 215; Rhesus monkeys have 1,623; and marmosets have 3,978, as detailed in an article published in the journal Genome Biology and Evolution in 2015.

RCPedia
The GAPDH gene that takes part in metabolizing carbohydrates: 50 retrocopiesRCPediaIn order for the information to be useful to other researchers, the two scientists set up the open access online database RCPedia, available since 2013. In 2014, a team from Adam Mickiewicz University of Poland released RetrogeneDB, a database that contains information about the retrocopies of 62 species, including insects, birds and amphibians.
Some of the differences between humans are now attributed to this mechanism for genome reformulation. In a study published in the journal PLOS Genetics in 2013, teams from Syrian-Lebanese, Indiana University, the AC Camargo Cancer Center and the Federal University of Rio Grande do Norte compared the genome of 17 individuals selected on the basis of the 1000 Genomes Project and concluded that there were an average of 91 retrocopies of 11 different genes between any two people; the genome of two people has 105 different genes on average. In a wider study, the team from São Paulo analyzed the genome of 2,557 people from 26 populations from all over the world and also detected variations among populations. Inhabitants of the interior of Africa presented unique retrocopies, different from those of Europeans.

Petr Kratochvil / Public domainGorilla: 215 unique retrocopiesPetr Kratochvil / Public domain
Independent of the parents
“Retrocopies represent a random and individual process of reformatting of the genome,” says Galante. Bolstering this conclusion in an article published in the journal Genome Research in March 2016, Francesco Carelli and other researchers at the University of Lausanne, Switzerland, contend that the retrocopies, although initially expressing in germinative cells – sperm and ova –, can establish themselves and diversify in somatic cells that make up living tissue. In addition, they can assume new functions, as complex as those of the parent genes from which they originated. In this way, the Swiss experts say, the parent genes could be lost with no harm to the organism they formed.
Retrocopies are generated directly by the messenger RNAs copied to an equivalent version of DNA that may establish itself in other genes, leading to the production of proteins different from those that were produced before its arrival. Geneticists recognize the intrusive gene because, unlike its original version and the genes common to the two, it does not contain segments that could lead to the formation of proteins, the so-called introns. Retrocopies were known as pseudo-genes precisely because they do not contain introns and do not always activate the process of producing proteins that build living creatures.
Retrocopies are now even viewed as “essential to the continuity of a species,” Galante argues. For example, he points to the PTEN gene that generates a protein capable of stopping tumor growth. The action of this gene can be reduced through molecules known as microRNAs, which adhere to it and reduce its action. PTENP1, a retrocopy of PTEN, attracts some of the microRNAs and thus, the PTEN can satisfy its function of impeding the growth of abnormal cells.
RCPEDIA
RPL21, the gene most often retrocopied in primates: nearly 200 copies in each speciesRCPEDIAThe team from Syrian-Lebanese is working on a similar case. “We have discovered a gene related to cell proliferation whose retrocopy functions the same way as PTENP1 does, reinforcing the hypothesis that the retrocopies constitute a mechanism of genomic regulation,” he says. The inverse of this can also be seen. In general, retrocopies are blocked by a mechanism known as methylation. If this block disappears and the retrocopies become active, they can alter the functioning of the genes in which they have become established and cause mutations that could favor the growth of tumor cells.
The hypothesis that they function as a safety system for important information helps us to understand the apparent exaggeration of having nearly 200 copies of the retrocopy known as RPL21, which makes up ribosomes, the structures responsible for the production of proteins essential to all humans, scattered among all the chromosomes.
Representatives of the group of retrocopies known as Alu are five times smaller than the common gene and abundant, having a million copies in the genome, equivalent to 11% of human DNA. Inversely, representatives of the family of retrocopies known as long interspersed nuclear elements-1 (LINE-1) are on average four times longer than common genes and comprise nearly 20% of the human genome. They are also distinct because they manage to act independently, without the need of RNAs, because they possess two segments that allow for their duplication.

RCPEDIA
The POLH gene for DNA repair: only a single copy. Mutations in the original gene are associated with xeroderma pigmentosumRCPEDIAIn 2010, Brazilian geneticists Alysson Muotri at the University of California in San Diego and Maria Carolina Marchetto at the Salk Institute in La Jolla introduced in the journal Nature the protein responsible for controlling the activity of LINE-1, abundantly expressed in stem cells and associated with the formation of neurons, and when deregulated, with mental disturbances such as schizophrenia and autism. Their work continues today. “We’re creating animal and human models in which these genes are silenced or over-expressed to see how they affect the nerve networks and behavior,” Muotri explains. According to a study by the University of Utah published in the journal Mobile DNA in May 2016, the 124 mutations already identified on LINE-1 could result in genetic diseases.
Experts in this type of jumping gene are anxiously awaiting the release of data from large international projects that are comparing genes expressed in various tissues – one of them with samples from 450 people, another from 1,500 people – in the hopes of gaining clarity on DNA segments once viewed as parasites or innocuous.
Project
Retrocopies: origins, polymorphisms and somatic variations (nº 2012/24731-1); Grant Mechanism Young Investigators in Emerging Markets; Principal Investigator Pedro Alexandre Favoretto Galante (Teaching and Research Institute of the Syrian-Lebanese Hospital); Investment R$843,619.40.
Scientific articles
CARELLI, F.N. et al. The life history of retrocopies illuminates the evolution of new mammalian genes. Genome Research. V. 26, No. 3, p. 301-14. 2016.
NAVARRO, F.C.P. and GALANTE, P.A.F. A genome-wide landscape of retrocopies in primate genomes. Genome Biology and Evolution. V. 7, p. 2,265-75. 2015.
ROBBIANI, F. D. et al. Plasmodium infection promotes genomic instability and AID-dependent B cell lymphoma. Cell. V. 162, No.4, p. 727-7. 2015.
SCHRIDER, D.R. et al. Gene copy-number polymorphism caused by retrotransposition in humans. PLOS Genetics. V. 9, No. 1, e1003242. 2013.
MUOTRI, A. et al. L1 retrotransposition in neurons is modulated by MeCP2. Nature. V. 468, No. 7,322, p. 443–6. 2010.