 FABIO OTUBOLittle more than a decade ago, only a few complete genomes were available for analysis. Today, there are not enough programs or personnel to track the number of DNA sequences that are deposited in public databases and produced every day by the new generation of sequencers. These extremely fast machines identify the base pairs, or chemical letters, of the genetic material at a cost that is thousands of times lower than was possible in the early 2000s, when the epic journey of sequencing the first human genome came to an end. Eyeing that challenge, mathematician João Meidanis, a founding partner of the company Scylla Bioinformática and professor at the University of Campinas (Unicamp) in Brazil, invested in a line of research to create simpler, more efficient methods of comparing two or more genomes.
FABIO OTUBOLittle more than a decade ago, only a few complete genomes were available for analysis. Today, there are not enough programs or personnel to track the number of DNA sequences that are deposited in public databases and produced every day by the new generation of sequencers. These extremely fast machines identify the base pairs, or chemical letters, of the genetic material at a cost that is thousands of times lower than was possible in the early 2000s, when the epic journey of sequencing the first human genome came to an end. Eyeing that challenge, mathematician João Meidanis, a founding partner of the company Scylla Bioinformática and professor at the University of Campinas (Unicamp) in Brazil, invested in a line of research to create simpler, more efficient methods of comparing two or more genomes.
Working with his former doctoral student Pedro Feijão in 2009, he formulated the theoretical basis for a method of comparing entire genomes, known as the Single-Cut-or-Join (SCJ) operation, and last year, he tested it in practice on the genomes of organisms, including plants and bacteria. “With our method, we can easily compare two or more genomes without exponentially increasing the number of calculations we make, which is what happens with other techniques,” Meidanis says. “We can use it to construct genealogical trees and see which genomes are closest or farthest from an evolutionary standpoint.” The mathematician was one of the bioinformatics coordinators for the project that, in 2000, sequenced the genome of the bacteria Xylella fastidiosa, which causes citrus variegated chlorosis in orange trees. The work resulted in the first cover story of the scientific journal Nature devoted to a Brazilian research study.
To compare all of the genetic material of one species to that of another, researchers must resort to simplification. The primary way to do this is to take into account the notion that the genes in the compared genomes are exactly the same but are in a different order in the specific sequence of each organism. Using this logic, methods for comparing genomes count the number of rearrangements that would be needed to transform one genome into another. These rearrangements occur when large segments of DNA in the original sequence move over time. The fewer rearrangements separating two genomes, the closer they are to one another on the evolutionary tree.
 Using their method, Meidanis and Feijão formulated an alternative definition for the concept of the breakpoint, an important parameter for finding rearrangements in a sequence and calculating the proximity of two genomes. A breakpoint is the location at which there is an interruption in a long conserved segment in the genomes being compared.
Using their method, Meidanis and Feijão formulated an alternative definition for the concept of the breakpoint, an important parameter for finding rearrangements in a sequence and calculating the proximity of two genomes. A breakpoint is the location at which there is an interruption in a long conserved segment in the genomes being compared.
Last year, the two researchers refined another method of genome comparison that is more elaborate than SCJ. This second technique, initially proposed in 2000, compares only circular genomes. With this development, it also became useful to compare the genetic material of linear chromosomes. “That was one of the limitations of the original technique,” says Feijão, who is now with Scylla. The new method, based on what mathematicians call adjacency algebraic formalism, has not yet been tested on real genomes. For now, it exists only in theory.
Metagenomics
Meidanis is clearly not the only researcher to feel the effects of the new reality in his field. Having returned to Brazil in mid-2011, after eight years at the Virginia Bioinformatics Institute in the US, João Carlos Setubal, who now is a full professor at the Chemistry Institute of the University of São Paulo (USP), notes that the demand for services and research in his field has grown in volume and sophistication in the recent years. As an example of this trend, he has received 16 proposals for collaboration on other researchers’ initiatives since returning to São Paulo. “The latest-generation sequencers produce an astronomical amount of data on genomes, proteomes and organism metabolism,” says Setubal, who was a bioinformatics coordinator for the Xylella project. “Because of declining technology costs, today any research project with minimal resources can sequence the genome of an organism.”
One field that has opened up to biologists and bioinformaticians in the past decade is metagenomics, which studies the microbiota of an ecological niche. Setubal’s principal research, an FAPESP thematic project on microorganisms at the São Paulo Zoo, is focused on this field. In this approach, instead of isolating and cultivating the microorganisms such that the DNA of each species can be extracted separately, he takes a sample directly from the environment to be studied. In such a sample, the DNA of several species comes “mixed,” and it is up to the bioinformaticians to find the techniques for separating and characterizing the genetic material of each species. “We are studying three microbiomes at the Zoo: compost made by zoo staff, water from the lakes and manure from howler monkeys,” Setubal says.
Metagenomics is also a way to search for unknown organisms in a specific habitat. The team headed by Ana Tereza Ribeiro de Vasconcelos, the coordinator of the bioinformatics center at the National Scientific Computing Laboratory (LNCC) in the city of Petropolis, participated in the discovery of magnetic bacteria found in Araruama Lake on the coast of the state of Rio de Janeiro, one of the world’s most saline lagoons. One of the bacteria found in that study was Candidatus magnetoglobus multicellularis; identified by Ulysses Lins of the Federal University of Rio de Janeiro (UFRJ), the bacteria are difficult to isolate from the environment and keep in a culture medium. “We are currently involved in about ten metagenomics projects,” says Vasconcelos, who has three sequencers in her laboratory and a team of approximately 25 people.
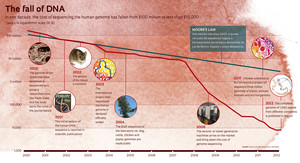
The amount of time and money required for projects devoted to DNA analysis of organisms has changed radically in the past decade. In the early years of the genomics era, only large companies dared to venture into this new field. By the time the international public consortium that sequenced human genome for the first time was officially terminated in April 2003, that mega-initiative required 13 years of work by hundreds of scientists from at least 18 countries, including Brazil, and had an estimated cost of $2.7 billion. In considerably lower but equally massive proportion, the sequencing of the Xylella bacterium had cost FAPESP $12 million and involved the contributions of 192 researchers over a three-year period.
Genome sequencing has become cheaper by a factor of 10,000 to 20,000 compared to what it cost a little more than a decade ago, according to the data from the National Human Genome Research Institute (NHGRI) in the U.S. The mass influx of the second-generation sequencers into the market in the early 2008, which used a technology different from that of the early Sanger-type machines, caused the cost of sequencing to plummet at a rate that far outpaced the performance gains resulting from Moore’s Law of computing power. Today, in two or three days and at a cost of just a few thousand dollars, it is possible to identify all of the three billion chemical letters of a person’s DNA. “Bioinformatics is a new tool, a magnifying glass, that enables us to better understand this biological phenomenon, which has not changed but can now be seen in another way,” says Gonçalo Pereira of the Institute of Biology (IB) at Unicamp.
However, sequencing is one thing, and extracting useful information from the billions of data points that computers pour out on a daily basis into the hands of scientists is another thing and is considerably more complex. “Genome sequencing is cheap today and has become a commodity, but data analysis is expensive,” says computer scientist João Paulo Kitajima of Mendelics, a new company that provides personalized genome analysis. “The number of people searching for jobs in bioinformatics has grown exponentially, and there is a supply and demand gap for specialists in Brazil and elsewhere.”

EBENGTSO / WIKIMEDIA COMMONSThe hypersaline waters of Araruama Lake in the state of Rio de Janeiro, where metagenomic studies found magnetic bacteriaEBENGTSO / WIKIMEDIA COMMONS
It is difficult to accurately estimate the size of the community of bioinformaticians in Brazil. According to Guillherme Oliveira, president of the Brazilian Association for Bioinformatics and Computational Biology (AB3C), there are approximately 300 people, including professors, students and researchers, who maintain ties with the organization. “Bioinformaticians were once self-taught,” says Oliveira, who coordinates the bioinformatics center at the Oswaldo Cruz Foundation (Fiocruz) in the state of Minas Gerais. “Today, many of them have come out of post-graduate programs, and every state has a bioinformatics specialist. What’s new is that now companies are also operating in this field.” Large Brazilian universities, such as USP, UFRJ and the Federal University of Minas Gerais (UFMG), as well as Fiocruz, have specific post-graduate programs in bioinformatics. Other universities incorporate it as a line of research in their post-doctoral programs in broader fields, such as biology or computing.
The work of sequencing and analyzing the genome of Schistosoma mansoni, the parasite that causes schistosomiasis, has been the focus of the highest profile project at the bioinformatics center of the Fiocruz facility in Minas Gerais in recent years. However, the six sequencing machines and 15 specialists in the bioinformatics unit headed by Oliveira have participated in approximately 60 different projects, including studies of the genomes of cancer, infectious agents, bovine breeds, and metagenomics studies. The center now also generates and analyzes data for the Research Network for the Molecular Identification of Brazilian Biodiversity (BR-BoL), coordinated by Cláudio Oliveira of the Institute of Biosciences at the Universidade Estadual Paulista (Unesp) in Botucatu, which plans to catalogue 120,000 specimens from 24,000 species in nature within four years. BR-BoL is the Brazilian arm of the International Barcode of Life Project, whose goal is to identify species by characterizing their DNA.
Bioinformatics has spread throughout Brazil, even to centers far from its major cities in the Southeast. At the Federal University of Pará (UFPA), Artur Silva conducts bioinformatics research in collaboration with groups from São Paulo. Since May of 2012, Sandro de Souza, who for years headed research in this field at the Ludwig Institute for Cancer Research in São Paulo, is now at the Brain Institute at the Federal University of Rio Grande do Norte (UFRN). He does not have a sequencer at his own facility in Natal, the state capital, but he appears unconcerned. “Don’t forget that I can even do sequencing in the cloud if I want,” Souza says. “I’m beginning my neuroscience research without any problem.”
However, Souza also has access to all of the machines from the Ludwig Institute, which closed its facility in the city of São Paulo and moved them to the Ribeirão Preto School of Medicine at USP (FMRP-USP), where the Center for Medical Genomics opened just last year. “The techniques used in genomics and bioinformatics will create a revolution in medical practice similar to what happened with image-based medicine,” says Wilson Araújo da Silva Júnior, one of the people in charge of the new center at FMRP.
To increase access to the DNA and RNA sequencing and analytical services, Unicamp is opening the Central Laboratory for High-Performance Technologies (LacTAD) on March 1. The laboratory will focus on genomics, proteomics, cell biology and bioinformatics. Its equipment includes two new-generation sequencers made by Illumina that are capable of decoding a complete human DNA sequence in a matter of a few days and a third machine to sequence specific genomic regions. The machines at the center have been in use since last year, when they arrived at the university, but were scattered around in different locations. Next month they will begin operating in the 2,000-m2 facility built for LacTAD.
“We believe there is an unmet demand for this type of service, and bioinformatics has become a bottleneck for many biological research studies,” says Ronaldo Pilli, a chemist and Provost for Research at Unicamp, who heads the project in the new laboratory. “We are joining the worldwide trend towards offering this type of service on a centralized basis, which makes it easier to purchase, operate, and update the machines.” LacTAD’s equipment was purchased for about R$5.5 million through FAPESP’s Multiuser Equipment Program. The building, budgeted at R$4 million, was financed by the university.
LacTAD will provide services to researchers at Unicamp and to other universities and businesses. Interested researchers can obtain price quotes for using the services at the laboratory website. “The cost of the work we do will range from R$100 to R$100,000,” Pilli says. Democratization has come to the world of bioinformatics.

Public domain image A giant panda: One of the genomes for which China’s Beijing Genomics Institute (BGI) decoded the complete sequencePublic domain image
China has the largest sequencing center
In less than 15 years, a Chinese bioinformatics center has gone
from being a minor partner in the international consortium that mapped the first human genome to a major global power in DNA sequencing. Established in 1999, the Beijing Genomics Institute (BGI) today has 180 sequencing machines, most
of which are latest generation units that can produce six terabytes
of data per day, an equivalent
of the complete genomes of 2,000 individuals. The center has 4,000 employees and affiliates in the United States, Europe and Japan.
This operation, conducted by
the Chinese on an enormous scale, has created expectations that the
cost of sequencing of a human genome will soon fall to $1,000.
Their work makes them a major
player in the state-of-the-art
projects that reach far beyond decoding the genetic sequence
of the giant panda, the national symbol, three years ago.
In 2010, for example, BGI sequenced the first complete genome of a human ancestor from the DNA of an Eskimo who lived 4,000 years ago. In 2012, it provided the DNA of 100 Chinese for the international effort to study the genome of approximately 1,000 people from different regions around the globe. In addition, last year, the center announced plans to sequence three million genomes of humans, plants, animals and microorganisms in the next few years.
The Chinese policy is an aggressive one both scientifically and commercially. Beyond selling its bioinformatics services, BGI is trying to ensure its own access to the most recent advances in the field. Early this year, the center received the go-ahead from the U.S. for the $177 million purchase of Complete Genomics, a California company that developed a new sequencing method. The results obtained using this method have been reported to be more accurate than those obtained with the current methods used worldwide.
Projects
1. Studies on the microbial diversity of the São Paulo zoo (nº 11/50870-6); Grant mechanism BIOTA Program – Thematic project; Coordinator João Setubal (USP); Investment R$1,711,698.25 (FAPESP)
2. EMU: Central Laboratory for High-Performance Technologies (nº 09/54129-9); Grant mechanism Multi-user Equipment Program; Coordinator Fernando Ferreira Costa (Unicamp); Investment R$ 6,034,431.00 (FAPESP).
