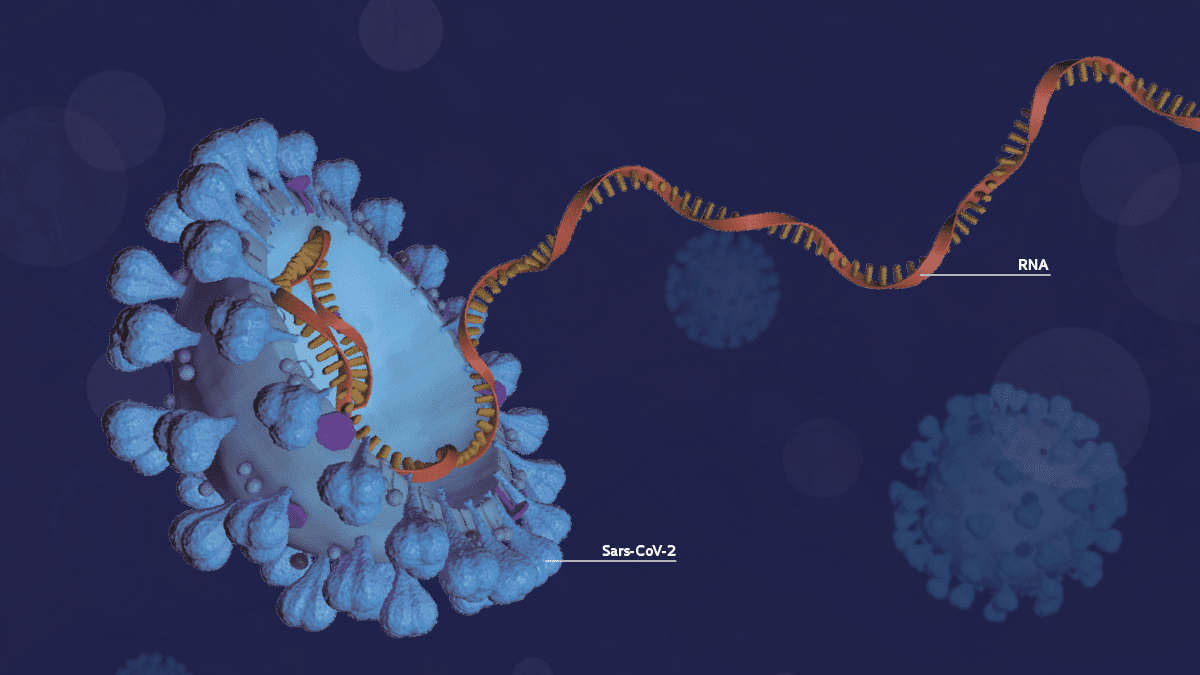
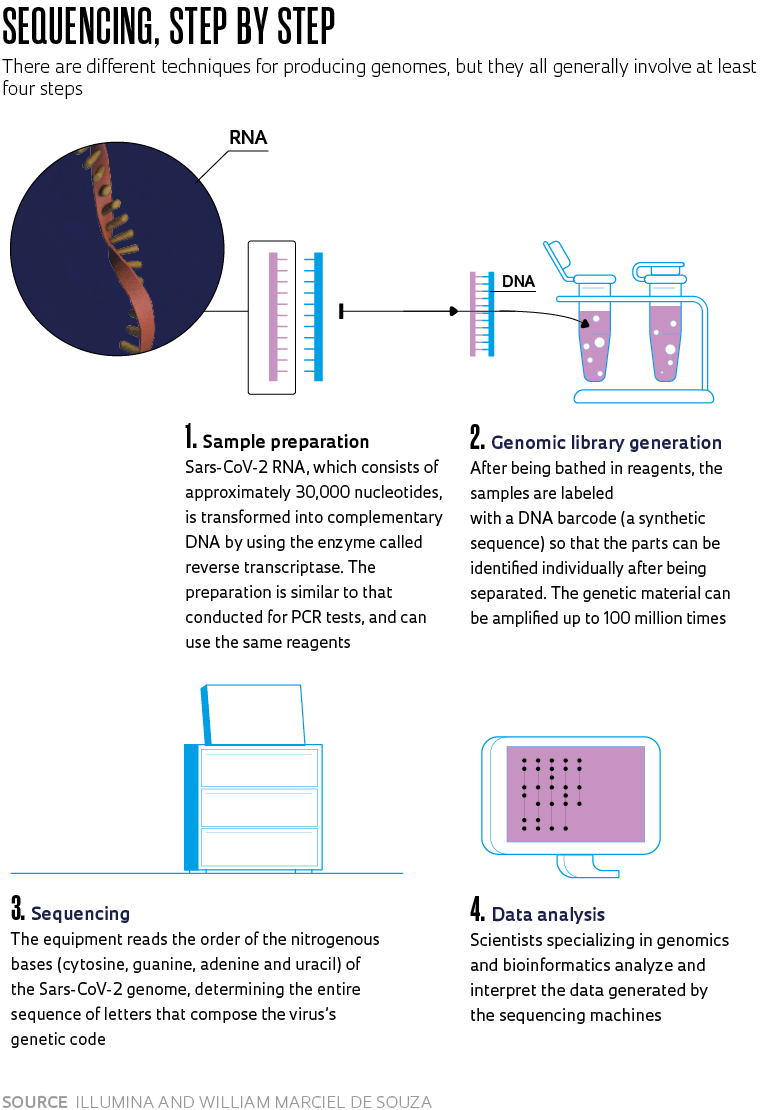
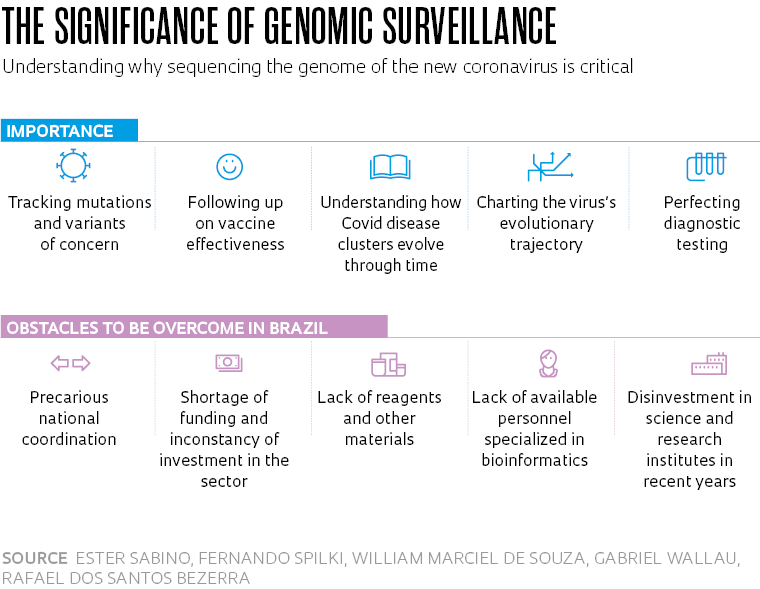
Understanding the genome of the Sars-Cov-2 virus present in infected populations is essential to tracking mutations and mapping the strains now circulating in Brazil. It’s also important in understanding how clusters of Covid-19 (the disease caused by the virus) evolve through time, and key to ensuring the accuracy of diagnostic testing and the effectiveness of vaccines and reconstructing the pathogen’s phylogeny (it’s evolutionary trajectory). However, Brazil’s genomic surveillance of Sars-CoV-2 is still far behind where it should be 15 months after confirming its first case of Covid-19. Despite the efforts of researchers working in the field, only 0.059% of the nearly 15 million positive new coronavirus cases recorded in Brazil as of May 20 had had their genomes sequenced and deposited on the GISAID platform, which is the principal international database for sharing data on the genetic code and mutations detected in the pathogen.
“The 5,000 samples sequenced so far are a tiny number for an entire year of epidemic,” says Dr. Ester Sabino, coordinator of the Joint Brazil-UK Centre for Arbovirus Discovery, Diagnosis, Genomics, and Epidemiology (CADDE), funded by FAPESP and the Medical Research Council of Britain. Sabino, a professor at the Institute of Tropical Medicine (IMT) at the School of Medicine of the University of São Paulo (FM-USP), made this statement during a public hearing held in April by the External Commission to Combat Covid-19 of the Brazilian House of Representatives.
At that time, GISAID data showed that Brazil was responsible for depositing 5,488 out of the 1,050,224 complete genomes shared on the platform since January 2020, equal to 0.52% of the sequencing conducted worldwide. The number increased to 9,316 in the second half of May, but the proportion in relation to the world total changed little and represents an extremely small percentage considering that the country accounted for 9.6% of the world’s confirmed cases of Covid-19 and 12.6% of total deaths.
In absolute numbers, 1.58 million Covid-19 genomes had already been sequenced by 172 countries as of May 20, 2021. The United States appeared in first place, with 451,000 genomes shared, surpassing the United Kingdom (408,000), which had maintained the lead for some time. As a proportion of the total recorded Covid-19 infections in a given country, the ranking was led by Iceland, which sequenced 77.7% of positive cases, Australia (59.1%) and New Zealand (46.4%).
The importance of carrying out genomic surveillance saw increased focus in Brazil in January, after the alarming P1 variant was identified in Manaus. The new strain spread quickly throughout the city, most likely beginning in November 2020, even though an estimated 67% of the population had been exposed to Sars-CoV-2 during the first wave of the pandemic in April and May 2020. Scientists estimate that P1 is between 1.7 and 2.4 times more transmissible than variants identified so far in the region and may be unaffected by immunity formed through previous infection.
Over a period of approximately seven weeks the proportion of new coronavirus samples classified as P1 in São Paulo went from zero to 87%, according to Sabino and her colleagues in an article published in the journal Science on April 14 of this year. The study also confirmed, as the researchers suspected, that the second outbreak in Manaus is associated with the emergence and rapid spread of P1, now considered a new variant of concern (VOC). This is the designation given by the World Health Organization (WHO) and national health authorities to strains that pose a greater potential risk to public health, either because they are more transmissible or virulent, or because they present new disease symptoms.
P1 is part of a group of variants of highest concern to governments and scientists worldwide during the first months of 2021. The others are B.1.1.7, first detected in Great Britain, B.1.351, identified in South Africa, and B.1.617, which contributed to the explosion of Covid-19 cases in India (see Pesquisa FAPESP issue no. 302) and was also classified as a VOC. In her speech before the Brazilian House, Sabino highlighted that without an expansion of genomic surveillance in Brazil, it will not be possible to confront the new variants that are certain to emerge. The main obstacles to such an increase, the researcher argues, are the absence of a nationally coordinated response to the epidemic and the lack of resources for sequencing.
In Brazil, two national networks interconnect institutions and researchers in an effort to decode the Sars-CoV-2 genome, identifying the strains present in different genetic states and mutations. The FIOCRUZ Genomic Network is supported by the Ministry of Health and coordinated by scientists from the Oswaldo Cruz Foundation (FIOCRUZ). The Covid-19 National Omics Network (Rede Corona-ômica BR) receives resources from the Ministry of Science, Technology and Innovation (MCTI) and brings together researchers from institutions such as USP, the University of Campinas (UNICAMP), and the National Laboratory of Scientific Computing (LNCC) in Rio de Janeiro, among others.
In the state of São Paulo, the Sars-CoV-2 Emerging Variants Pandemic Alert Network was established in March, coordinated by the Butantan Institute, with the participation of teams from USP, São Paulo State University (UNESP), São José do Rio Preto School of Medicine (FAMERP) and Mendelics, a private company that specializes in genetic analysis. In a sample collected in March in the city of Sorocaba, the network’s researchers detected the presence of the South African variant B.1.351 in Brazil for the first time. “We confirmed it twice, by the mutations and through its phylogenetic tree,” said biomedical and bioinformatics specialist Rafael dos Santos Bezerra, from the Hemocentro de Ribeirão Preto. He is one of the authors of an article, still awaiting peer review, which describes detecting the B.1.351 strain through genomic monitoring.
Participants in these networks expect to scale up both the volume of sequencing and the country’s capacity to conduct genomic surveillance over the next few months. “The Rede Corona-ômica labs received large batches of processing materials at the end of March. We expect a big jump in sequencing over the coming weeks,” says virologist Fernando Spilki, the network’s coordinator and a professor at Feevale University, in Nova Hamburgo, Rio Grande do Sul. Spilki adds that the new processing materials are enough to produce about a thousand genomes, almost a fifth of what was deposited during the first year of the pandemic.
The processing materials, in this case, are kits with the necessary reagents to prepare the samples and “run” the sequencing machines. The kits are mostly imported from the United States and Europe, as are the sequencers, many of which had already been brought to Brazil due to the Zika virus outbreak in 2015. Few companies manufacture the equipment and reagent kits. One of the biggest is the US firm Illumina, whose sequencing systems have a reputation for reliable results. Another is British manufacturer Oxford Nanopore Technologies, which markets its portable MinION sequencer.
“All of the research groups are facing difficulties in receiving sufficient reagents to enable them to respond at scale,” says biologist Gabriel Wallau, from Instituto Aggeu Magalhães, of FIOCRUZ, in Recife. “Since the United States ramped up its sequencing and the rest of the world is doing the same, we need to make a pretty convincing argument to the company to get the reagents.”
After assuming the US presidency earlier this year, Joe Biden made genomic surveillance a priority. During the first few months of 2021, the amount of sequencing more than doubled. Even so, in May only 1.5% of the positive Covid-19 cases were sequenced in the United States, below the 5% proposed—in a preprint article published on the medRxiv platform—as being ideal for detecting emerging variants before they surpass 1% of the cases within a determined location.
“We have the know-how like any other country, and we have quality equipment, but there are two primary, interconnected problems: there’s a lack of resources for sequencing and, where they do exist, there’s a disconnect in obtaining reagents, which don’t always arrive in a timely manner,” Wallau says. The laboratory where he works in Pernambuco was finalizing the purchase of reagents in early May. “I believe that shortly we will have a much better view of how the new coronavirus is evolving in Brazil.”
The municipality of São Paulo will soon have a program to monitor its variants of concern with a project developed within the scope of CADDE, the binational project coordinated by Sabino. The idea behind the initiative, organized by FAPESP in an arrangement with municipal government, is to analyze around 380 Covid-positive samples per week, chosen at random, in proportion to the population of each city district. In part of the samples the virus genome will be sequenced completely; in another portion a PCR test that can identify the presence of the variants of highest concern will be performed. In the pilot program, conducted during the first week of March, 71.2% of the 73 samples tested from the municipality’s residents were from VOC’s. Of these, 64.4% were P1, the Manaus variant, and 6.8% were B.1.1.7, initially found in the United Kingdom.
In Brazil, many laboratories that perform genomic sequencing of the new coronavirus also conduct diagnoses of the disease via PCR, or they have arrangements with other public and private institutions that do PCR testing, such as the Central Public Health Laboratories (LACEN). These agreements facilitate access to the samples to be sequenced.
Complete genome sequencing is the gold standard for discovering mutations and new variants of interest and concern in Sars-CoV-2, but a new tool has appeared on the radar of Brazilian researchers: real-time PCR-based genotyping, the exam used in virus detection. “One of the protocols used today was created by immunologist Felipe Naveca’s team at FIOCRUZ Amazonas,” Spilki adds. Using the protocol they developed, the molecular test is able to discern if a positive sample for the new coronavirus belongs to an already recognized VOC. “Thus, we were able to evaluate not just hundreds, but thousands of virus sequences, practically everything that’s already circulating. And we get results on the dispersal of variants more quickly.”
Similar technology has been developed by the Integrative Biology Lab at the Institute of Biological Sciences of the Federal University of Minas Gerais (ICB-UFMG). The advantages of this new method for detecting variants are its low price and speedy results, according to the team led by geneticist Renato Santana. The test costs an average of R$70 and is finished in four hours, while fully sequencing a virus sample can cost R$500 or more and take weeks to complete.
Wallau says another significant bottleneck for expanding genomic surveillance work in Brazil is the shortage of bioinformatics specialists. “It’s one thing to sequence, it’s another to analyze the sequence. More refined analyses take time,” the researcher observes. “We have very powerful machines that generate a huge volume of data, but how do we analyze all that data accurately?”
Paulo Chapchap, chairman of the board of directors of the United for Health Institute (ITpS), agrees. “We have some important bioinformatics centers dedicated to infectious diseases in Brazil, but we need to train more professionals,” he says. Supported by the Itaú Unibanco Foundation, ITpS has R$200 million to invest in initiatives aimed at genomic surveillance.
Cuts to investment in science over recent years, both in basic and applied research, have also affected investigations. “There was a severe disinvestment, which harmed universities and research centers, not just the federal government,” says biomedical physician William Marciel de Souza, a postdoctoral intern at Oxford University, in England, who has collaborated with Brazilian research groups to analyze the genomes sequenced in Brazil. “What is being done now in terms of sequencing by FIOCRUZ, Rede Virus, and various researchers around the country is a miracle. It is a fundamental effort to try to understand the pandemic in Brazil and help get it under control.”
Projects
1. Joint Brazil-UK Center for Arbovirus Discovery, Diagnosis, Genomics, and Epidemiology (CADDE) (nº 18/14389-0); Grant Mechanism Thematic Project; Principal Investigator Ester Cerdeira Sabino (USP); Investment R$5,271,740.75.
2. Assessment of the impact of emerging and re-emerging viruses in hemotherapy and stem cell transplantation using advanced molecular techniques (nº 17/23205-8); Grant Mechanism Young Investigator Award; Supervisor Svetoslav Nanev Slavov (USP); Investment R$1,534,184.25.
Republish